Publication-Quality Plots
This guide accompanies Genomics Exchange Session 4 (March 10, 2026). Building on the tidyverse data wrangling skills from Session 3, we now turn that clean data into publication-ready figures using ggplot2.
We will create five common genomics figure types — volcano plots, MA plots, heatmaps, enrichment bar charts, and multi-condition bar plots — using the same maize drought RNA-seq dataset from Session 3. Each section shows the default output first, then walks through the changes needed to make it ready for a journal.
-
Prerequisites
- Completed Session 3 (or comfortable with tidyverse basics)
- Access to RStudio on Negishi via Open OnDemand
- R packages:
tidyverse,ggrepel,pheatmap,viridis,RColorBrewer,scales
-
What you will learn
- Build five publication-quality figure types in ggplot2
- Apply colorblind-safe palettes (viridis, RColorBrewer)
- Export figures at journal-required DPI and dimensions
- Transform an ugly default plot into a polished figure
- Set a reusable global theme for consistent styling
Setup¶
-
Launch RStudio on RCAC
Go to gateway.negishi.rcac.purdue.edu, select Interactive Apps > RStudio Server, choose your allocation, and request 1 core, 4 GB memory, and 2 hours. Once the session starts, click Connect to RStudio.
Warning
Always run RStudio through Open OnDemand on a compute node. Do not run R sessions directly on login nodes — they are shared resources and intensive computation will be terminated.
-
Download the data files
In the RStudio Terminal tab (not the R console), run:
Tip
$RCAC_SCRATCHpoints to the correct scratch directory in all clusters. On Gautschi it is set to/scratch/gautschi/${USER}and on Bell/scratch/bell/${USER}and so on. -
Set your working directory in R
Switch to the R console and run:
-
Set the cairo graphics device
On headless HPC sessions (no X11 display), R's default PNG device may fail. Set the cairo backend before loading any plotting packages:
Note
This is only needed on cluster compute nodes. If you are running RStudio through Open OnDemand, it is usually handled automatically, but setting it explicitly does no harm and prevents cryptic errors.
-
Install and load packages
Install these packages if you have not used them before (run once):
Then load them:
-
Set a global theme
Every ggplot starts with a default theme. Instead of adding theme customization to every plot, set it once:
base_size = 14sets a readable default font size. All text elements (axis labels, titles, legend) scale relative to this value.
Load and prepare data¶
We reuse the four Session 3 files and add one new enrichment results file.
Prepare the DESeq2 results the same way we did in Session 3 — classify genes by direction and compute the transformed p-value:
Define a color palette that we will reuse across plots. These colors are distinguishable under the three common forms of color vision deficiency:
Volcano plot¶
The volcano plot is the signature figure of differential expression analysis. The x-axis shows biological effect size (log2 fold change) and the y-axis shows statistical significance (-log10 adjusted p-value). Genes in the upper corners are both statistically significant and biologically meaningful.
Quick version¶
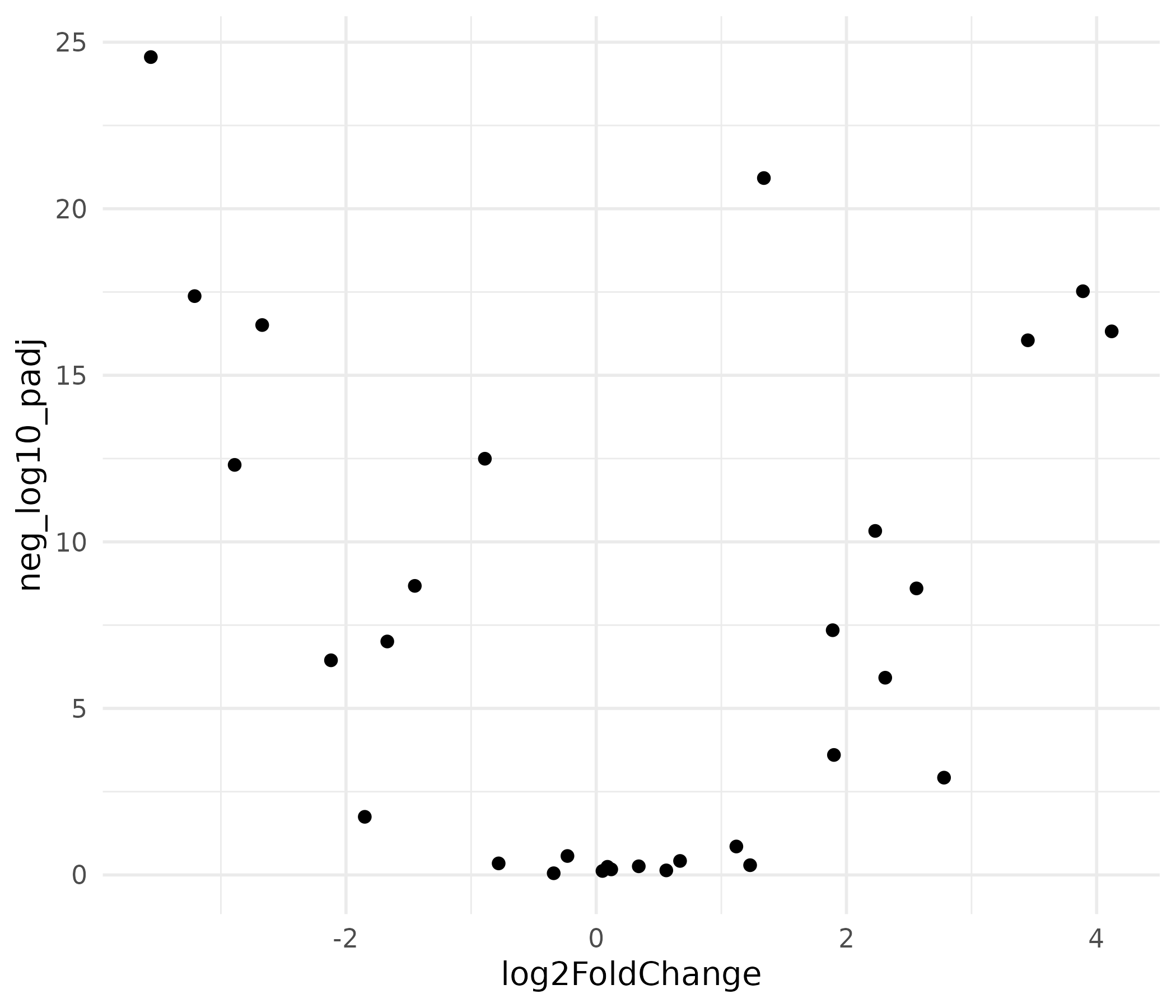
This produces a functional scatter plot, but it is not ready for publication. There is no color coding, no gene labels, no threshold lines, and the axis labels are raw variable names.
Publication version¶
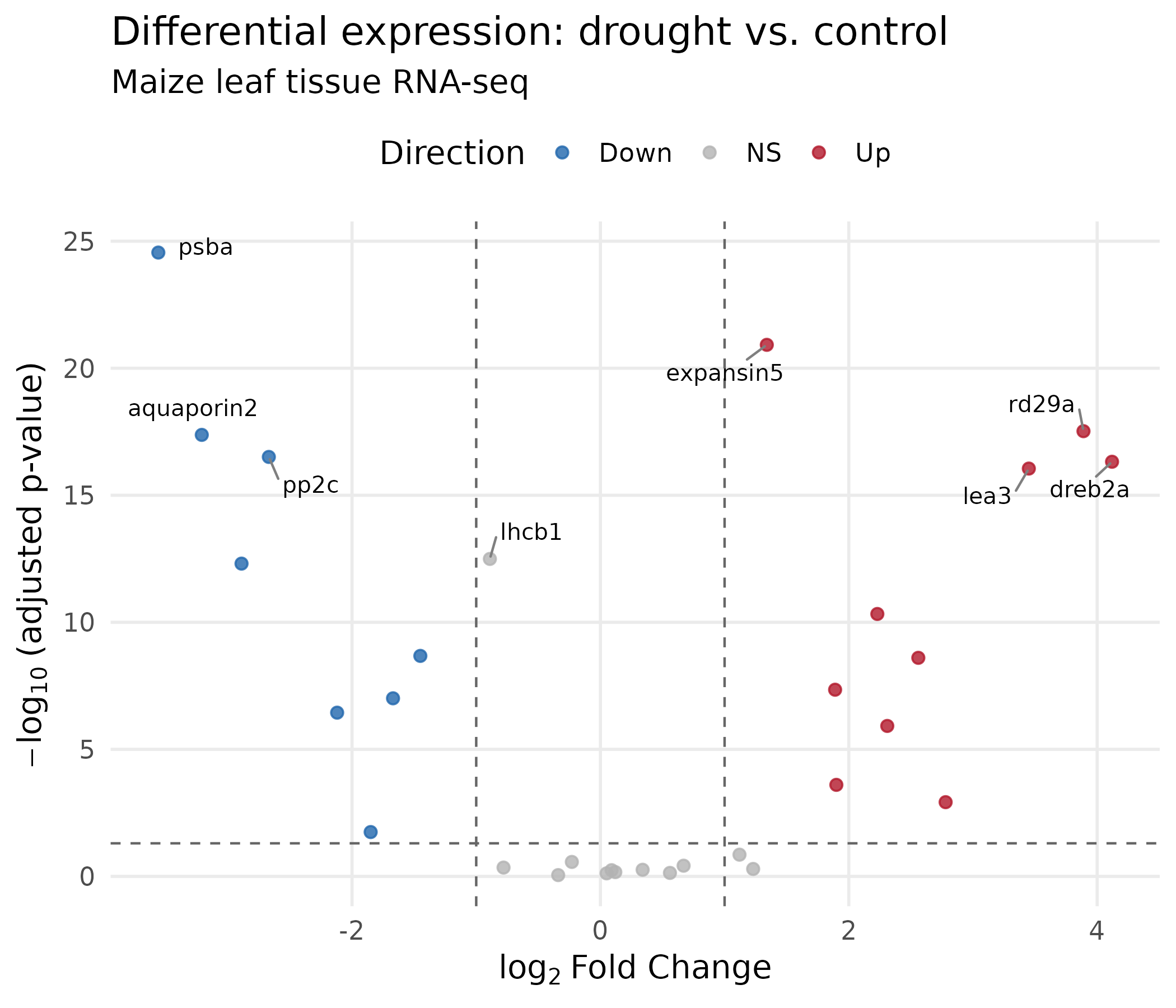
What changed and why:
geom_text_repel()from theggrepelpackage positions gene labels so they do not overlap. Thesegment.colorargument draws a line connecting each label to its point.- Dashed threshold lines at padj = 0.05 and |log2FC| = 1 show your significance cutoffs directly on the figure. Readers do not have to guess.
expression()renders mathematical notation:log₂with a proper subscript.scale_color_manual()maps direction to our colorblind-safe palette.
Export¶
Tip
Always save both PNG (for slides and web) and PDF (for journal submission). PDFs produce vector graphics that scale to any size without pixelation.
MA plot¶
The MA plot shows mean expression level (x-axis) vs. fold change (y-axis). It reveals whether fold change estimates depend on expression level — low-count genes tend to have noisier fold changes. This is a standard QC figure in RNA-seq analysis.
Quick version¶
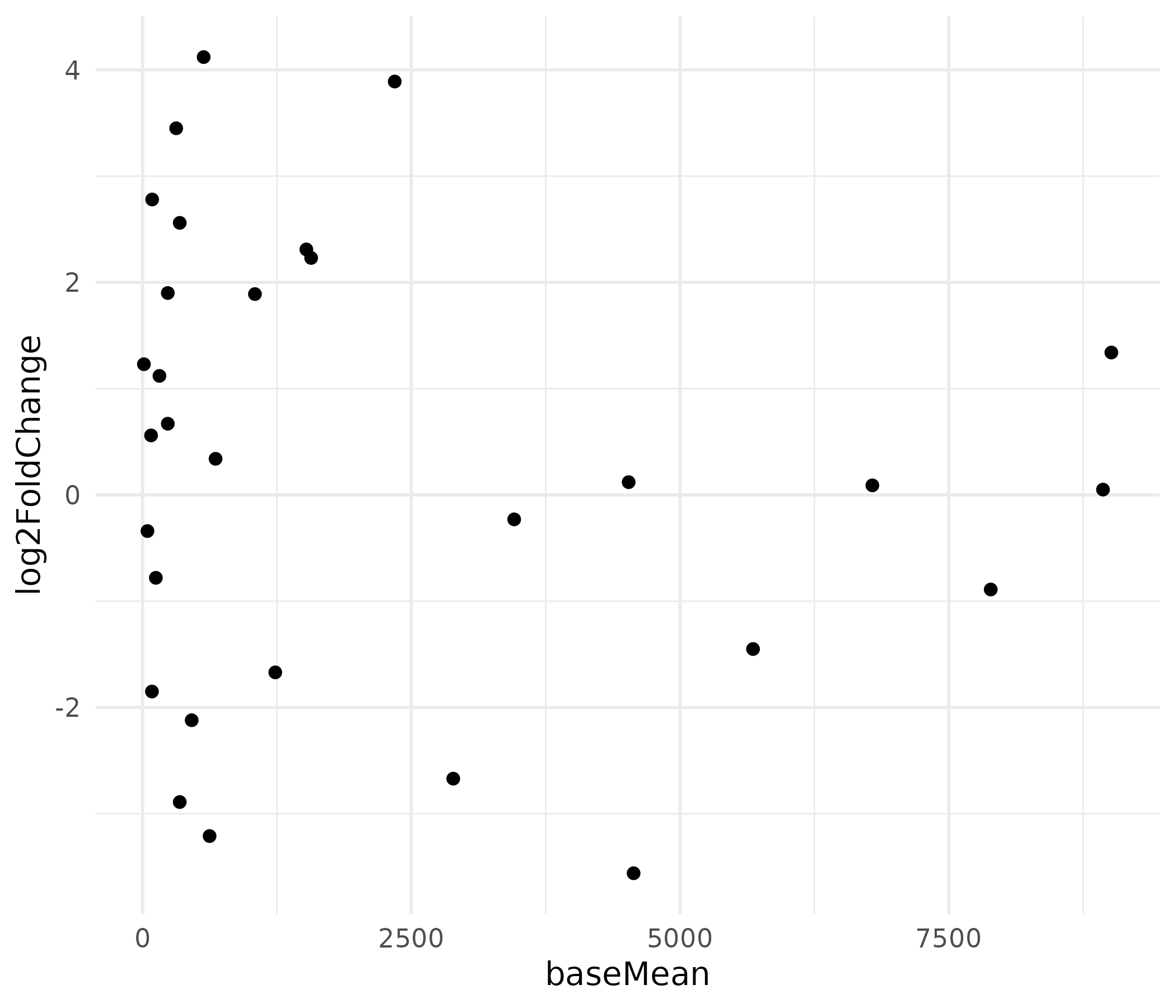
The x-axis is heavily right-skewed because a few highly expressed genes dominate the scale, compressing most points to the left.
Publication version¶
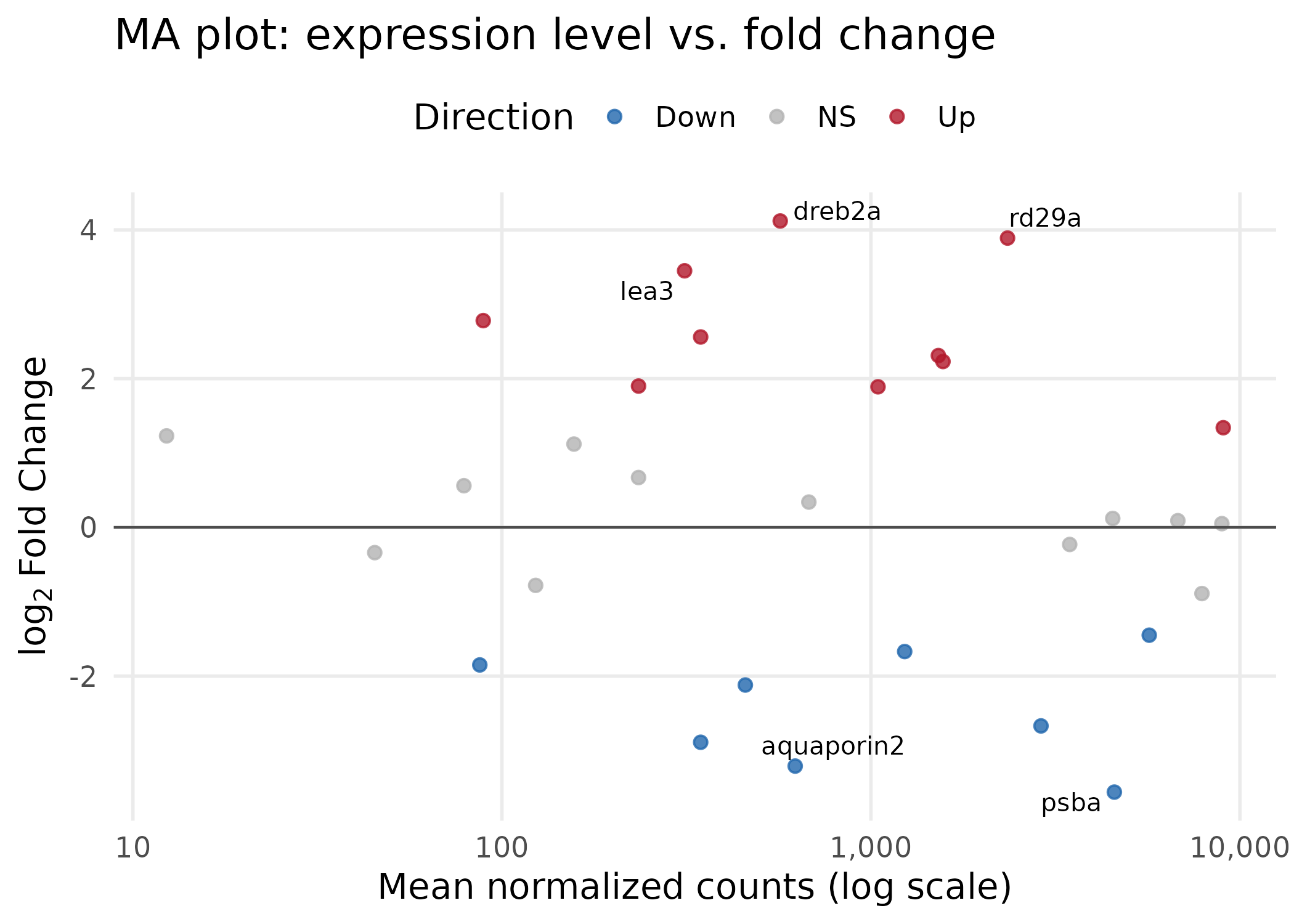
Key choices:
scale_x_log10()spreads the heavily right-skewed baseMean distribution, making all genes visible. Thelabel_comma()function from thescalespackage keeps axis labels human-readable (e.g., "1,000" instead of scientific notation).- The horizontal line at y = 0 provides a visual reference — genes above are upregulated, below are downregulated.
- Only genes with |log2FC| > 3 are labeled to avoid clutter.
Export¶
Heatmap¶
Heatmaps display expression patterns across genes (rows) and samples (columns) using color intensity. They are effective at showing groups of co-regulated genes and whether samples cluster by experimental condition. We use the pheatmap package, which handles clustering and annotation sidebars.
Prepare the data¶
Replace gene IDs with readable gene names:
Scale the rows (z-score normalization) so the color represents relative change within each gene, not absolute expression level. Without scaling, highly expressed housekeeping genes would dominate the color map:
Quick version¶
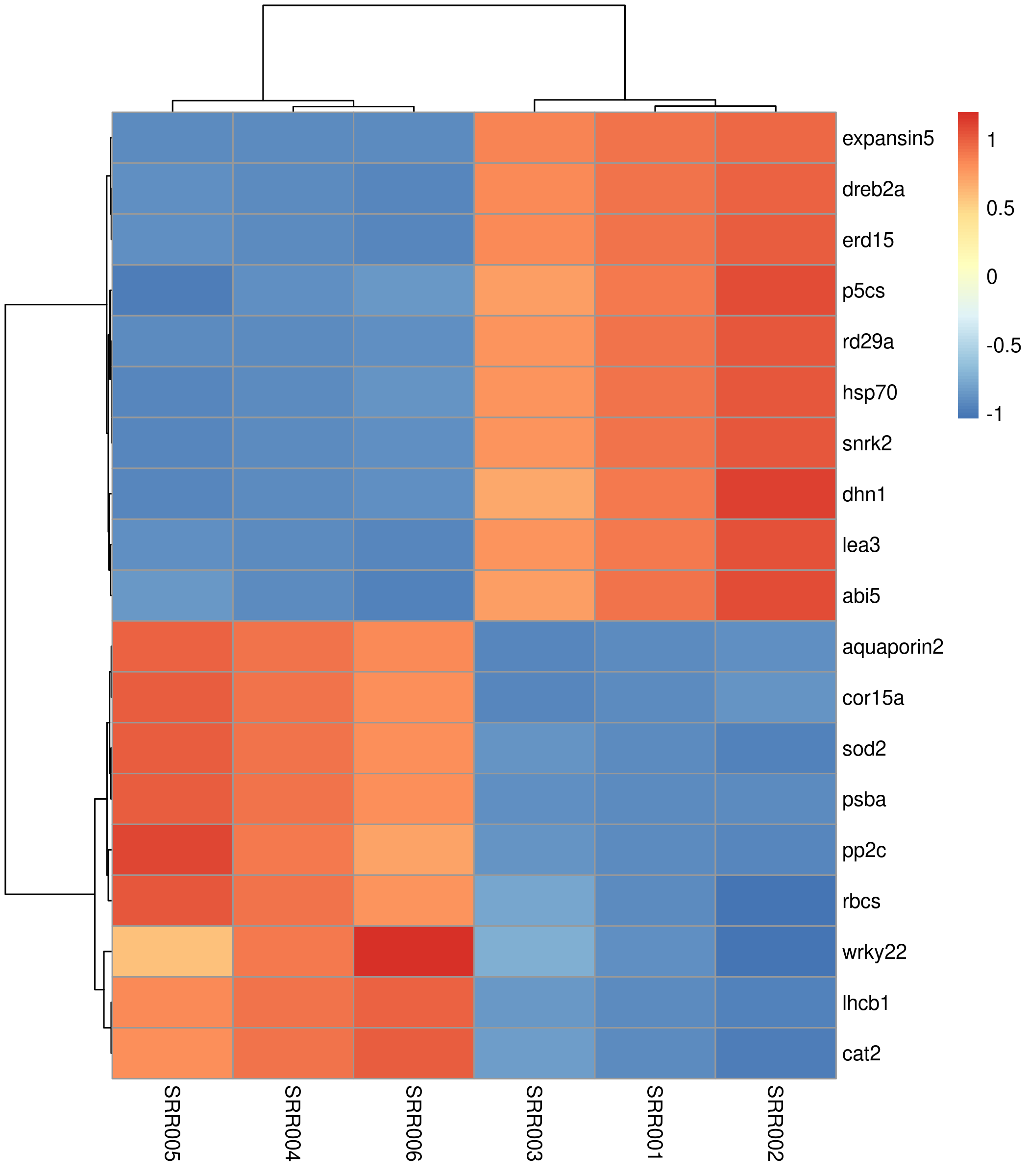
The default heatmap uses a blue-to-red palette, clusters both rows and columns, and works. But it lacks sample annotations and uses a suboptimal color scheme.
Publication version¶
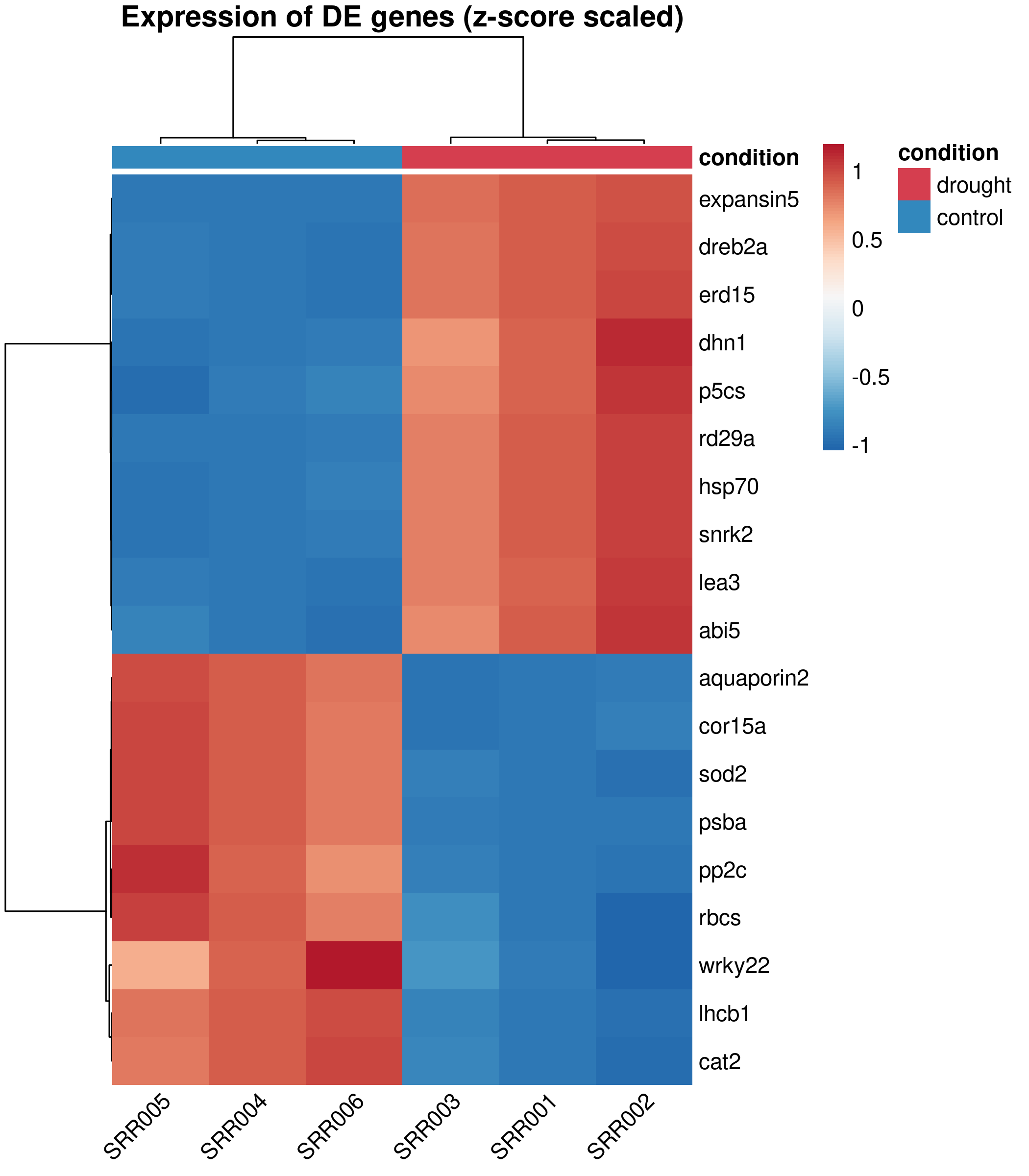
Design decisions:
RdBupalette (Red-Blue, diverging) is colorblind-safe and conventional for expression heatmaps. Blue = low, white = unchanged, red = high.annotation_coladds a colored sidebar showing which condition each sample belongs to, so readers immediately see the grouping.ward.D2clustering produces compact, well-separated clusters. It is the default recommendation for expression data.border_color = NAremoves the grid between cells, which adds visual noise in dense heatmaps.
Export¶
pheatmap does not return a ggplot object, so we use png() / pdf() with dev.off() instead of ggsave():
Warning
Do not forget dev.off() after png() or pdf(). If you skip it, the file will not be written and your RStudio plot pane may stop working until you run dev.off().
For PDF:
Enrichment bar plot¶
GO or functional enrichment analysis summarizes the biological themes among your differentially expressed genes. Here we use a small simulated enrichment result to demonstrate two common visualization styles: a dot plot and a bar chart.
The data¶
Each row is a GO term with its name, gene count, fold enrichment, and adjusted p-value.
Quick version¶
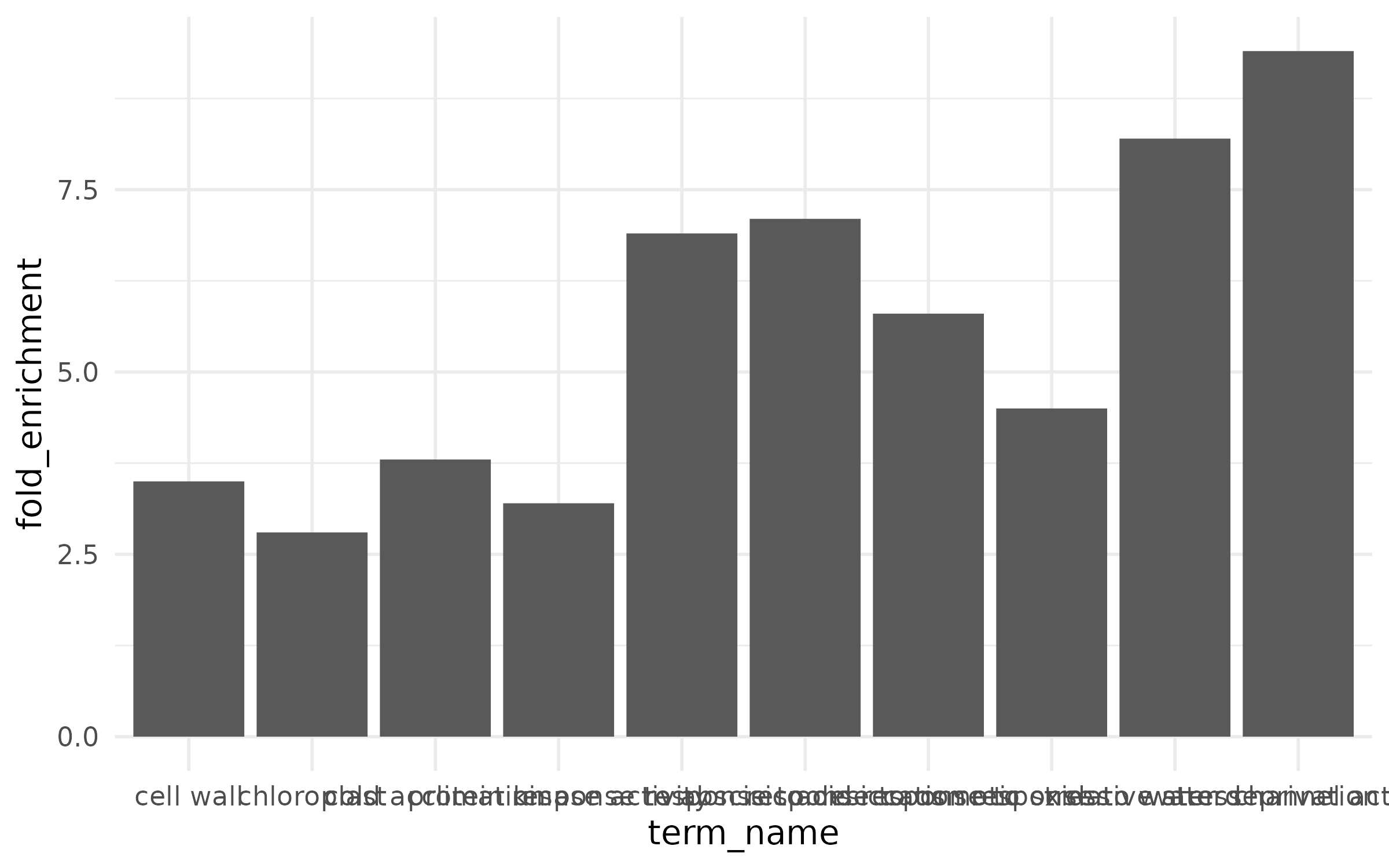
The x-axis labels overlap and are unreadable, there is no color to convey additional information, and terms are in alphabetical order instead of ranked by effect.
Publication version — dot plot¶
The dot plot encodes three variables: x-position (fold enrichment), color (category), and point size (gene count).
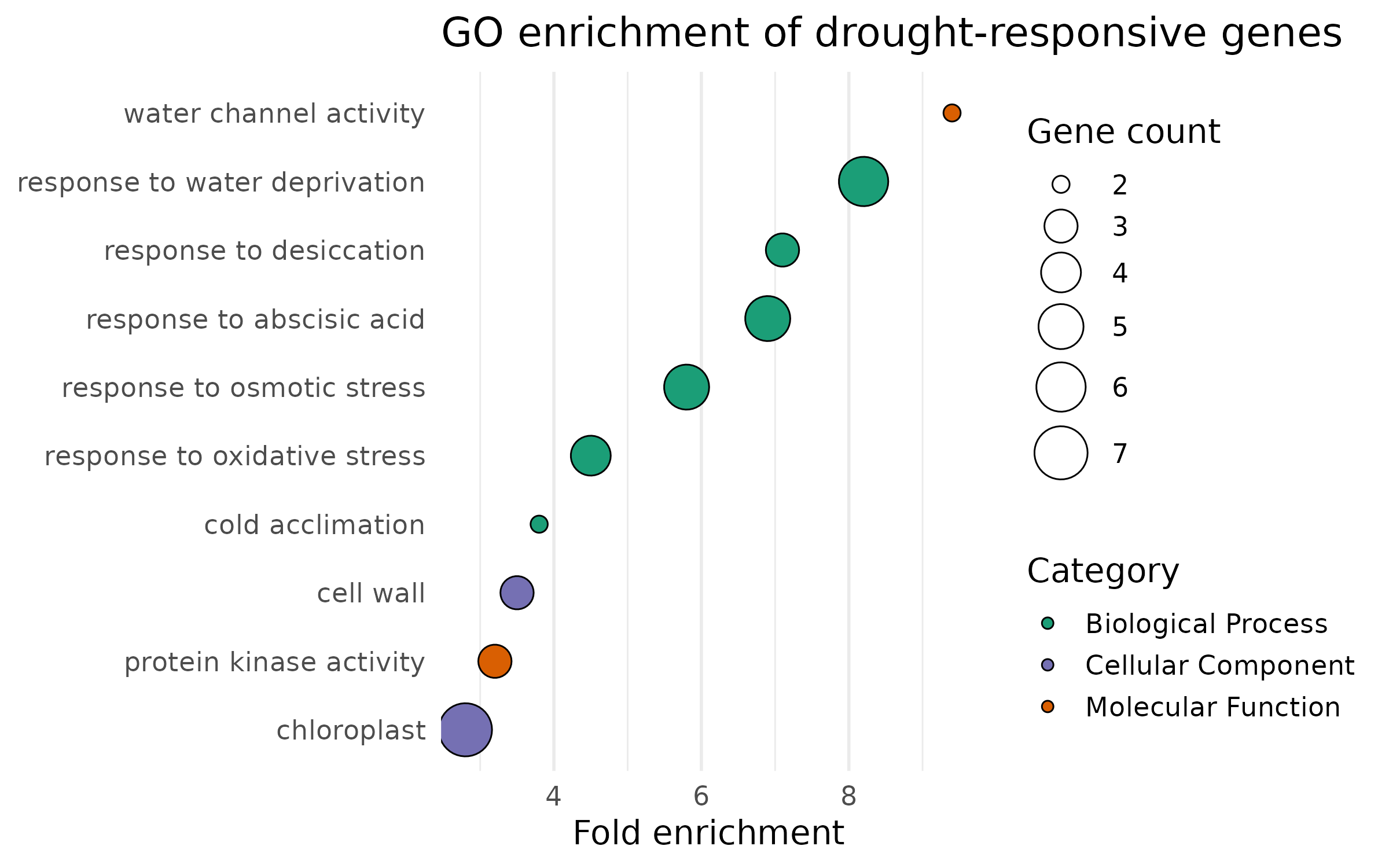
Publication version — bar chart¶
The bar chart uses color to encode significance instead of category:
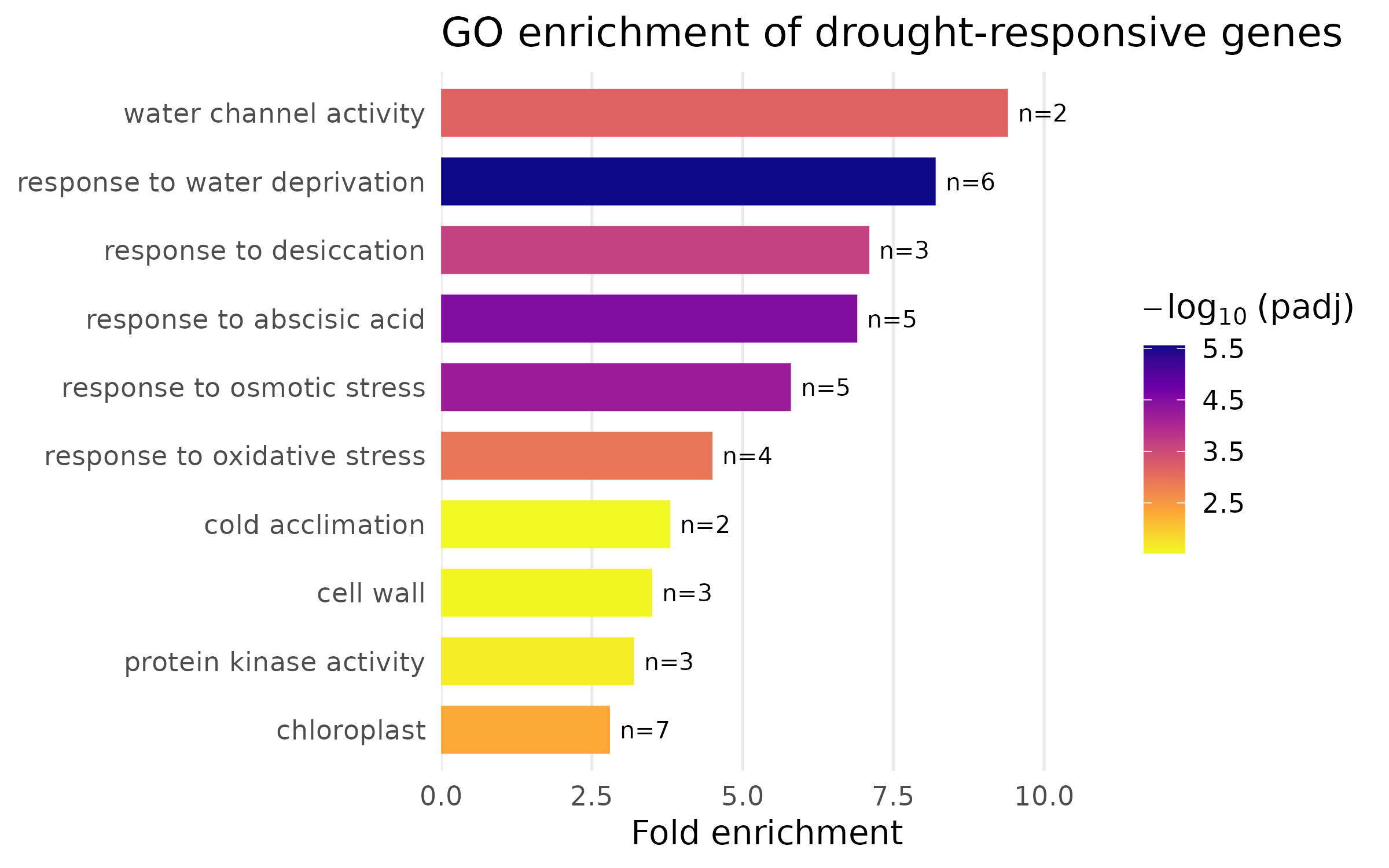
Design notes:
fct_reorder()sorts terms by fold enrichment so the most enriched terms appear at the top. Never leave categorical axes in alphabetical order — rank them by the variable of interest.- Horizontal layout (terms on the y-axis) avoids the overlapping-label problem entirely.
viridis"plasma" palette is perceptually uniform and colorblind-safe. The continuous color scale maps significance, giving the reader two dimensions of information.
Export¶
Before and after: ugly to publication-ready¶
This section demonstrates the iterative improvement process. We start with a default ggplot bar chart and fix one problem at a time.
Prepare data¶
Step 1: The ugly default¶
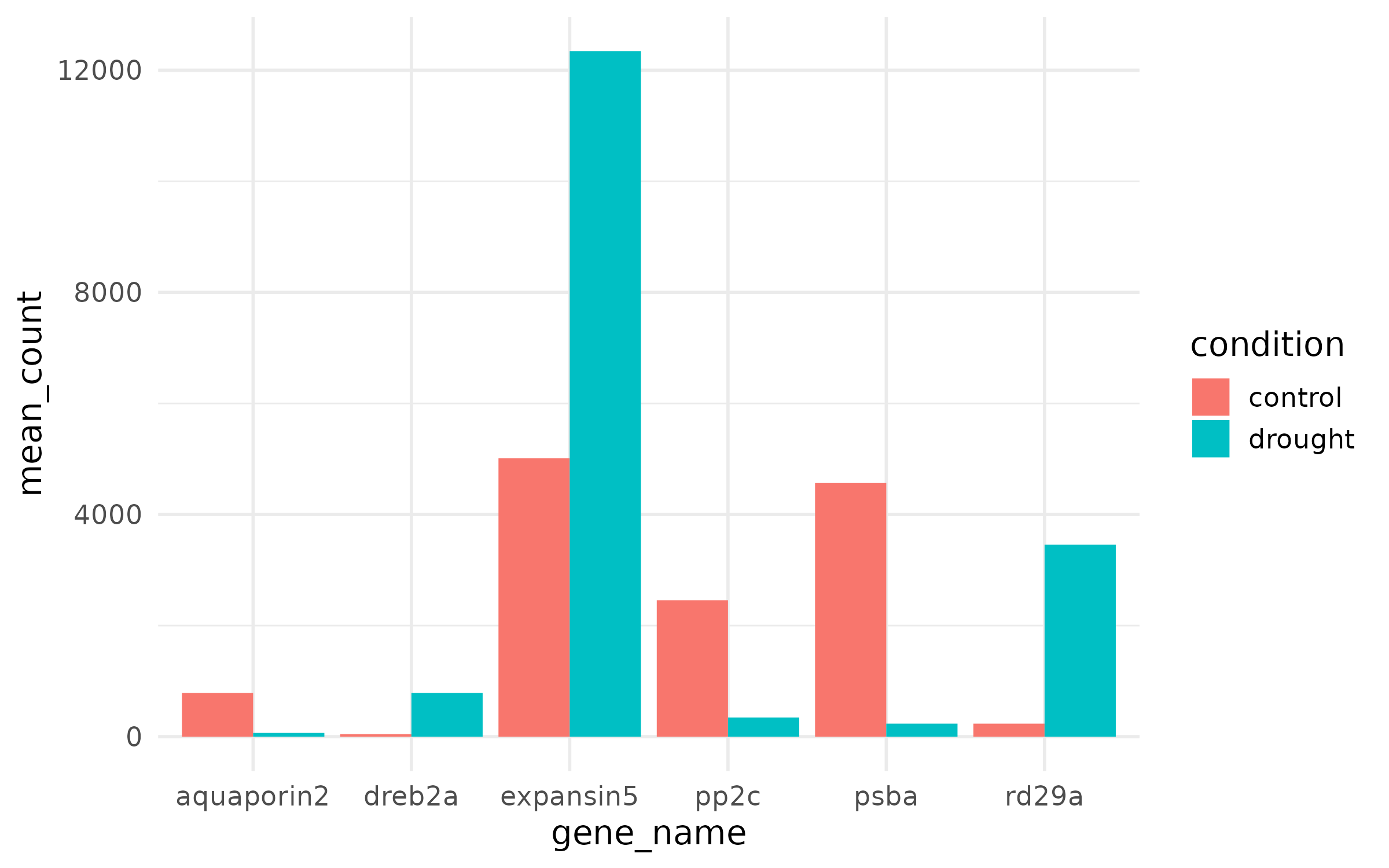
What is wrong:
- Default grey background is distracting
- Axis labels are raw variable names
- Default fill colors are not colorblind-safe
- No error bars — readers cannot assess variability
- Font is too small for a figure panel
- No title explaining what the figure shows
- Too many gridlines add visual noise
Step 2: Fix the theme¶
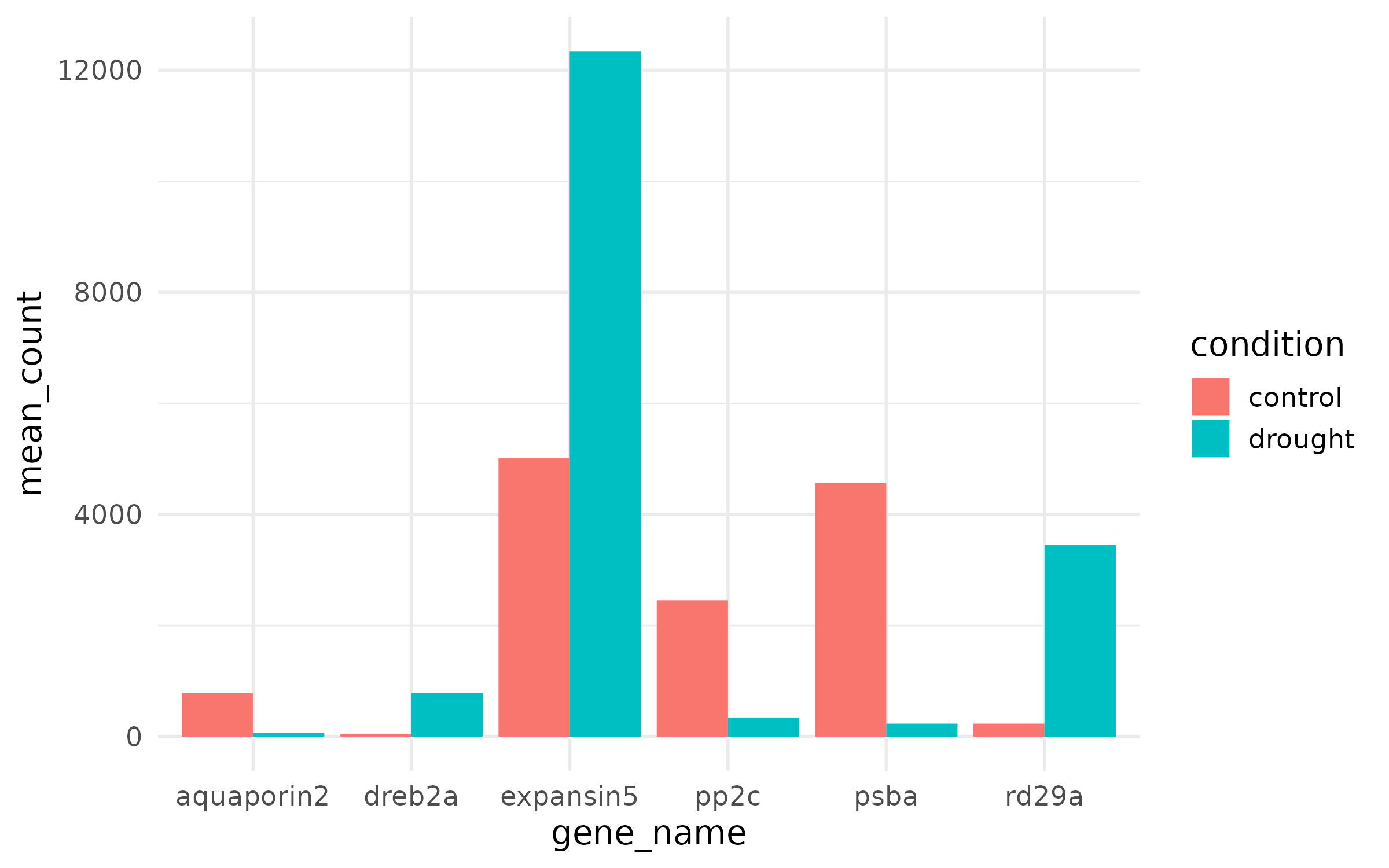
Step 3: Fix colors¶
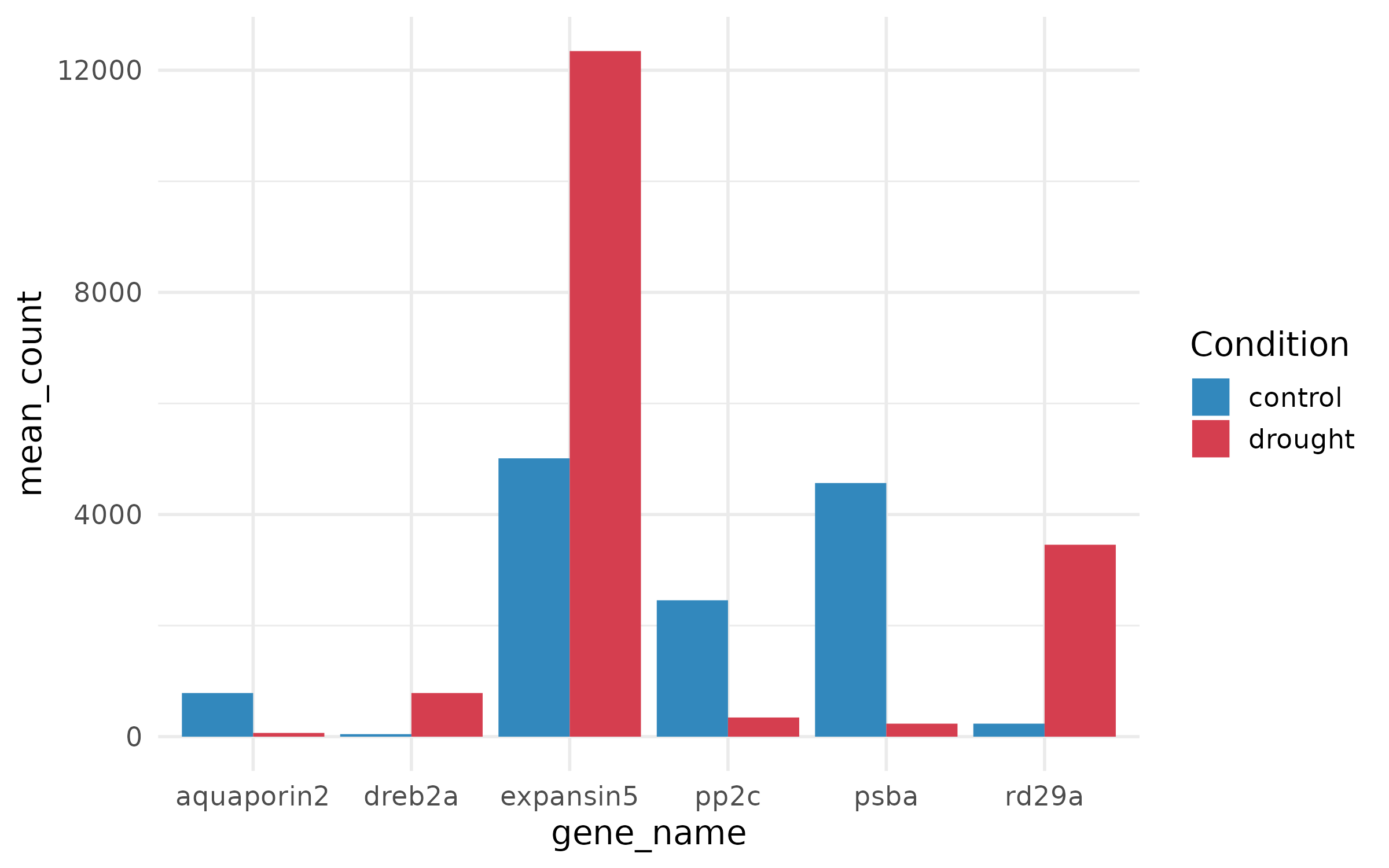
Step 4: Add error bars and proper labels¶
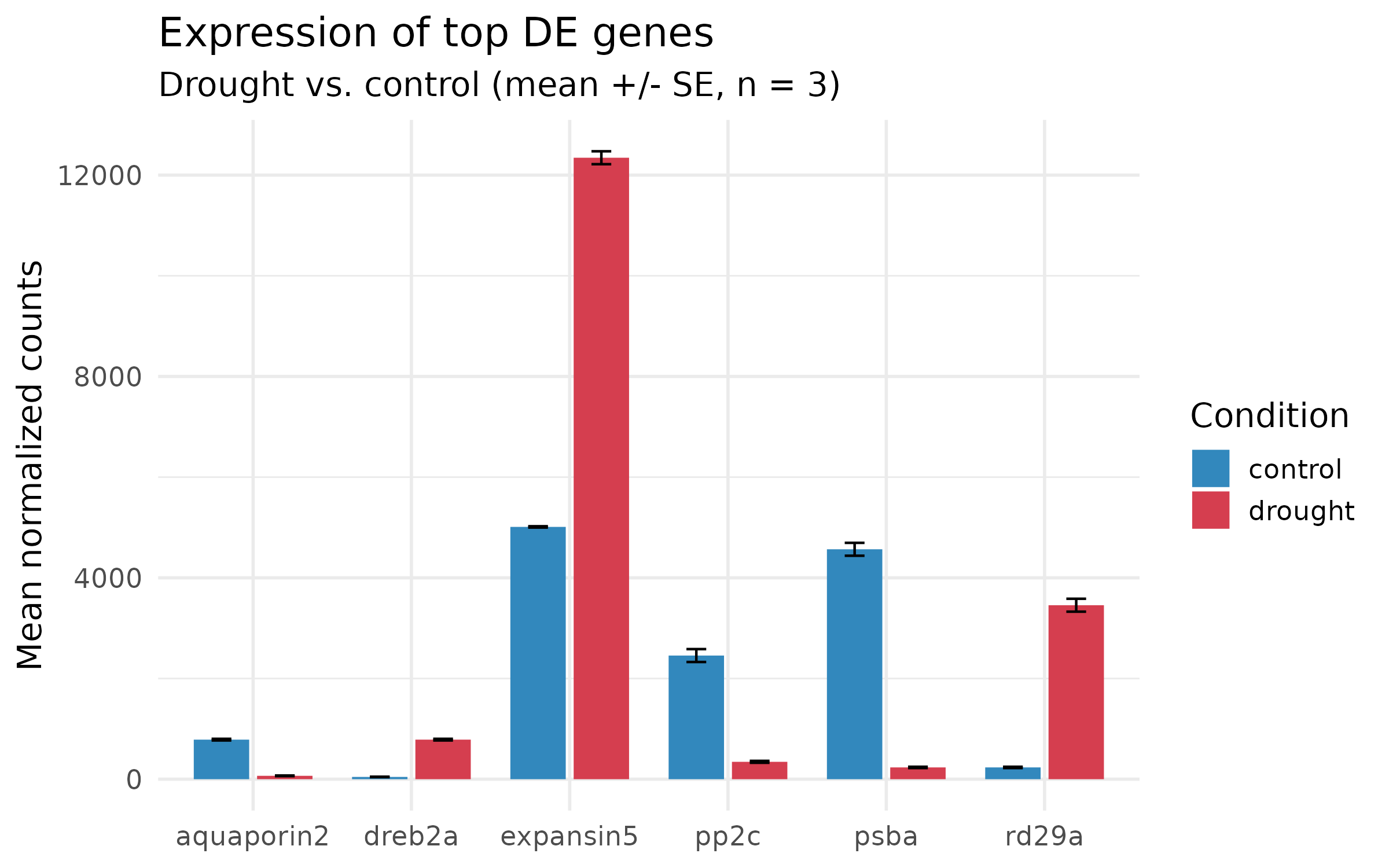
Step 5: Final polish — remove chartjunk¶
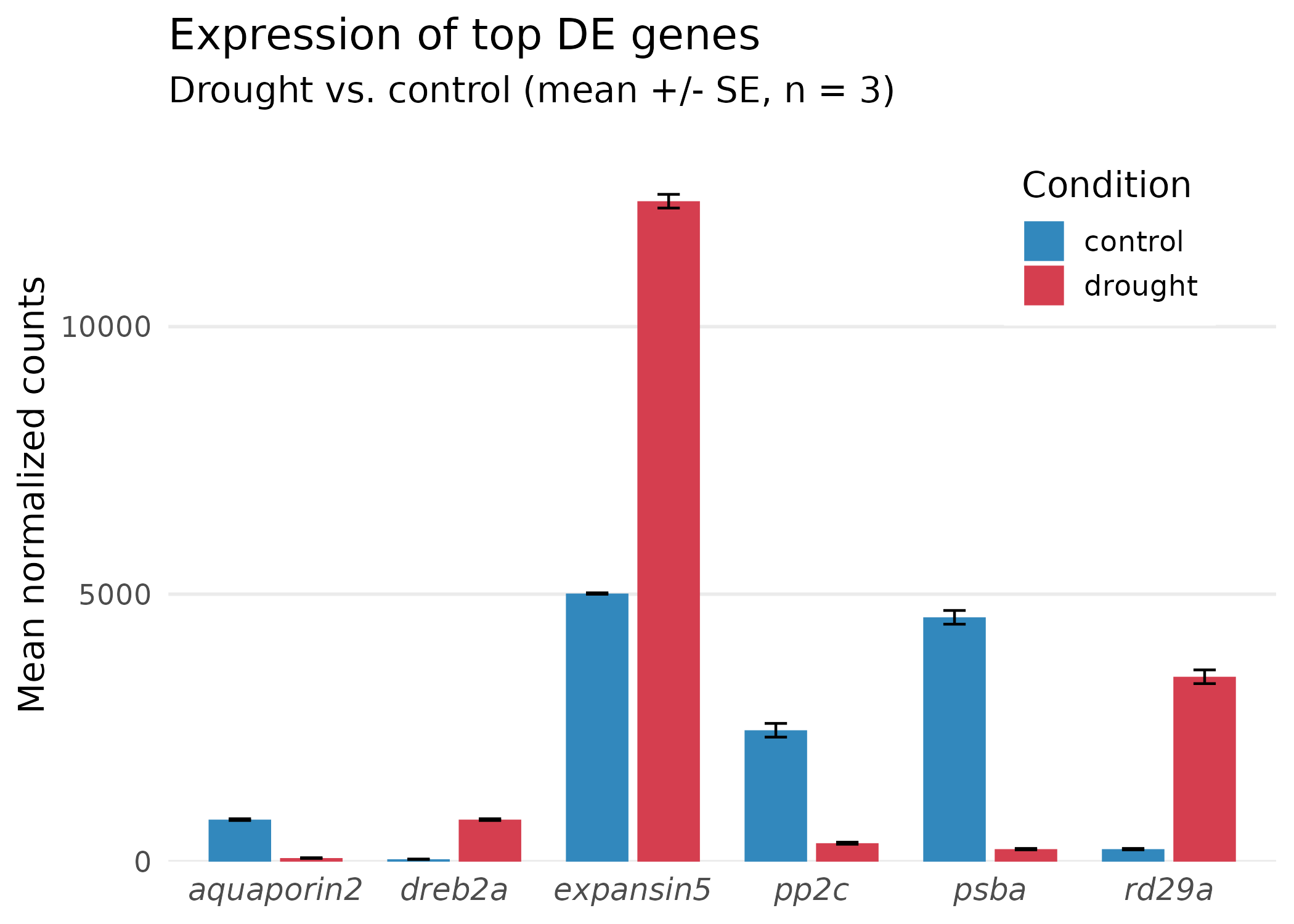
What changed in the final step:
- Removed vertical gridlines — they are not useful when the x-axis is categorical.
- Removed minor gridlines — reduces visual noise.
- Moved the legend inside the plot — saves space in tight figure panels.
- Italicized gene names — gene symbols are italicized by convention in biological publications.
- y-axis starts at zero — bar charts must start at zero to avoid misrepresenting magnitudes.
Export¶
Principles of publication figures¶
Color palettes for accessibility¶
Approximately 8% of men and 0.5% of women have some form of color vision deficiency. Every figure you publish should be readable by this audience.
Recommended palettes:
| Palette | Package | Best for | How to use |
|---|---|---|---|
| viridis | viridis |
Continuous data (expression, p-values) | scale_fill_viridis_c() |
| RdBu | RColorBrewer |
Diverging data (fold changes, z-scores) | scale_fill_distiller(palette = "RdBu") |
| Dark2 | RColorBrewer |
Categorical data (2-8 groups) | scale_color_brewer(palette = "Dark2") |
| Set2 | RColorBrewer |
Categorical data (lighter variant) | scale_fill_brewer(palette = "Set2") |
To see all colorblind-safe palettes in RColorBrewer:
Tip
When you only need 2–3 colors and want full control, define them manually with hex codes. The palette c("#2166AC", "#B2182B") (blue and red from the RdBu scale) works well for two-group comparisons and is safe for all common color vision deficiencies.
When to use which plot type¶
| Question | Plot type | Key ggplot2 function |
|---|---|---|
| Which genes are significant and large-effect? | Volcano plot | geom_point() + geom_text_repel() |
| Does fold change depend on expression level? | MA plot | geom_point() + scale_x_log10() |
| How do expression patterns cluster? | Heatmap | pheatmap() |
| What biological themes are enriched? | Enrichment dot/bar plot | geom_point() or geom_col() |
| How does one gene differ between conditions? | Box/bar plot | geom_boxplot() or geom_col() |
Journal figure requirements¶
Most journals specify figure dimensions and resolution. Common requirements:
| Journal tier | Format | DPI | Max width |
|---|---|---|---|
| Nature family | PDF or TIFF | 300 | 180 mm (7.1 in) |
| Cell family | PDF or EPS | 300 | 174 mm (6.85 in) |
| PLOS journals | TIFF or EPS | 300 | 174 mm (6.85 in) |
| General rule | PDF + PNG | 300 | 170 mm (~6.7 in) |
Standard column widths:
Always check your target journal's author guidelines before final export. Use ggsave() with explicit width, height, dpi, and units for every figure.
Font sizing rules of thumb¶
| Element | Minimum size | ggplot2 parameter |
|---|---|---|
| Axis labels | 8–10 pt | theme(axis.title = element_text(size = ...)) |
| Axis tick labels | 7–9 pt | theme(axis.text = element_text(size = ...)) |
| Legend text | 7–9 pt | theme(legend.text = element_text(size = ...)) |
| Title | 10–12 pt | theme(plot.title = element_text(size = ...)) |
| Panel labels (facets) | 8–10 pt | theme(strip.text = element_text(size = ...)) |
Note
When setting base_size in your theme (e.g., theme_minimal(base_size = 14)), all other text sizes scale relative to it. A base_size of 12–14 works well for most single-panel figures. For multi-panel figures that will be shrunk to fit a column, start with base_size = 10–12.
Reusable theme function¶
If you produce many figures for one project or paper, define a custom theme once and apply it everywhere:
ggplot2 cheat sheet¶
| Task | Function | Example |
|---|---|---|
| Scatter plot | geom_point() |
aes(x, y, color) |
| Bar chart | geom_col() |
aes(x, y, fill) |
| Error bars | geom_errorbar() |
aes(ymin, ymax) |
| Non-overlapping labels | geom_text_repel() |
aes(label) (from ggrepel) |
| Horizontal line | geom_hline() |
yintercept = 0 |
| Vertical line | geom_vline() |
xintercept = c(-1, 1) |
| Log scale | scale_x_log10() |
Spread skewed data |
| Manual colors | scale_color_manual() |
values = c("A" = "#hex") |
| Viridis continuous | scale_fill_viridis_c() |
Colorblind-safe gradient |
| Brewer discrete | scale_fill_brewer() |
palette = "Dark2" |
| Readable numbers | label_comma() |
10000 becomes 10,000 |
| Math notation | expression() |
expression(log[2]~"FC") |
| Reorder factor | fct_reorder() |
Sort categorical axis by a numeric variable |
| Save figure | ggsave() |
width, height, dpi, units |
| Set global theme | theme_set() |
Apply once, affects all plots |
Troubleshooting¶
ggsave produces a blank or corrupted PNG on the cluster
This is almost always a missing graphics device. Set the cairo backend at the top of your script:
If cairo is not available, try:
If it returns FALSE, use type = "Xlib" in ggsave() or export as PDF instead (which does not need a bitmap device).
ggrepel labels are missing or say 'too many overlaps'
Increase max.overlaps:
Or reduce the number of labeled points by filtering more strictly:
pheatmap colors look washed out or wrong
The number of colors in your palette must be large enough to represent the data range. Use at least 50–100 colors:
If your z-scores are extreme, the color scale may be dominated by outliers. You can set symmetric breaks:
My fonts look different in the saved file vs. the RStudio plot pane
The RStudio plot pane uses screen rendering, while ggsave() uses the file device. To get consistent results, always judge your figures from the saved file, not the plot pane. Open the PNG in an image viewer or the PDF in a PDF reader.
FAQs¶
Should I use pheatmap or ComplexHeatmap?
pheatmap is simpler and covers most use cases. ComplexHeatmap (Bioconductor) is more powerful — it supports multiple annotation tracks, split heatmaps, and oncoplots. If you need to show more than one annotation (e.g., condition + batch + genotype), switch to ComplexHeatmap. For a single condition annotation like we have here, pheatmap is sufficient.
Can I use these techniques with other organisms?
Yes. Everything here applies to any differential expression or enrichment result, regardless of organism. Swap the gene IDs, GO terms, and condition labels, and the same code works for human, Arabidopsis, mouse, or any species.
How do I arrange multiple plots into one figure?
Use the patchwork package:
This creates a 2-over-1 layout with panel labels A, B, C. Most journals require multi-panel figures as a single file.
What if my journal requires TIFF format?
The compression = "lzw" flag reduces file size significantly. Some journals accept LZW-compressed TIFFs; others require uncompressed. Check the author guidelines.
Next session¶
Session 5 (March 24): Running bioinformatics programs on RCAC. We will cover how to use the module system, run containerized tools with Apptainer, submit SLURM jobs, and manage bioinformatics workflows on RCAC clusters.
Resources¶
- Session 3 tutorial: R Skills for Biological Data — prerequisite data wrangling guide
- R for Data Science (2e) — chapters 2 and 10 cover ggplot2 fundamentals
- ggplot2 documentation — complete function reference
- Claus Wilke, Fundamentals of Data Visualization — free online book on visualization principles (not R-specific)
- ggplot2 cheat sheet (PDF) — print this
- ColorBrewer 2.0 — interactive palette explorer with colorblind simulation
- Genomics Exchange Discord — ask questions between sessions
- Questions? Email arun@purdue.edu